This paper is available on arxiv under CC 4.0 license.
Authors:
(1) Zheyu Oliver Wang, Department of Aeronautics and Astronautics, Massachusetts Institute of Technology, Cambridge, MA and olivrw@mit.edu;
(2) Ricardo Baptista, Computing + Mathematical Sciences, California Institute of Technology, Pasadena, CA and rsb@caltech.edu;
(3) Youssef Marzouk, Department of Aeronautics and Astronautics, Massachusetts Institute of Technology, Cambridge, MA and ymarz@mit.edu;
(4) Lars Ruthotto, Department of Mathematics, Emory University, Atlanta, GA and lruthotto@emory.edu;
(5) Deepanshu Verma, Department of Mathematics, Emory University, Atlanta, GA and deepanshu.verma@emory.edu.
Table of Links
- Abstract & Introduction
- Background and Related Work
- Partially Convex Potential Maps (PCP-Map) for Conditional OT
- Conditional OT flow (COT-Flow)
- Implementation and Experimental Setup
- Numerical Experiments
- Discussion and Reference
6. Numerical Experiments.
We test the accuracy, robustness, efficiency, and scalability of our approaches from sections 3 and 4 using three problem settings that lead to different challenges and benchmark methods for comparison. In subsection 6.1, we compare our proposed approaches to the Adaptive Transport Maps (ATM) approach developed in [4] on estimating the joint and conditional distributions of six UCI tabular datasets [29]. In subsection 6.2, we compare our approaches to a provably convergent approximate Bayesian computation (ABC) approach on accuracy and computational cost using the stochastic Lotka–Volterra model, which yields intractable likelihood. Using this dataset, we also compare PCP-Map’s and COT-Flow’s sampling efficiency for different settings. In subsection 6.3, we demonstrate the scalability of our approaches to higher-dimensional problems by comparing them to the flow-based neural posterior estimation (NPE) approach on an inference problem involving the 1D shallow water equations. To demonstrate the improvements of
Table 2
Hyperparameter search space for PCP-Map and COT-Flow. Here m denotes the size of the context feature or observation y

PCP-Map over the amortized CP-Flow approach in the repository associated with [20], we compare computational cost in subsection 6.4.
6.1. UCI Tabular Datasets.
We follow the experimental setup in [4] by first removing the discrete-valued features and one variable of every pair with a Pearson correlation coefficient greater than 0.98. We then partition the datasets into training, validation, and testing sets using an 8:1:1 split, followed by normalization. For the joint and conditional tasks, we set x to be the second half of the features and the last feature, respectively. The conditioning variable y is set to be the remaining features for both tasks.
To perform joint density estimation, we use the block-triangular generator h: R n+m → R n+m as in (2.2), which leads to
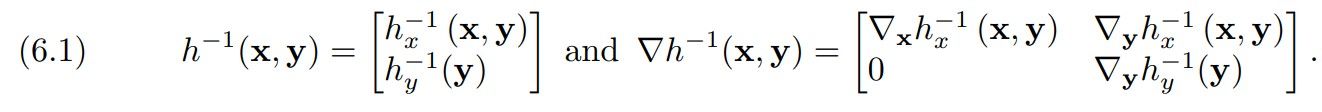
Here, the transformation in the first block, hx, is either PCP-Map or COT-Flow, and the transformation in the second block, hy, is their associated unconditional version. We learn the weights by minimizing the expected negative log-likelihood functional
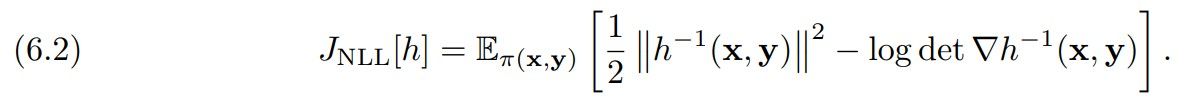
An alternative approach for joint density estimation is to learn a generator g where each component depends on variables x and y, i.e., g does not have the (block)-triangular structure in (6.1). This map, however, does not immediately provide a way to sample conditional distributions; for example, it requires performing variational inference to model the conditional distribution [54]. Instead, when the variables that will be used for conditioning are known in advance, learning a generator with the structure in (6.1) can be used to characterize both the joint distribution π(x, y) and the conditional π(x|y).
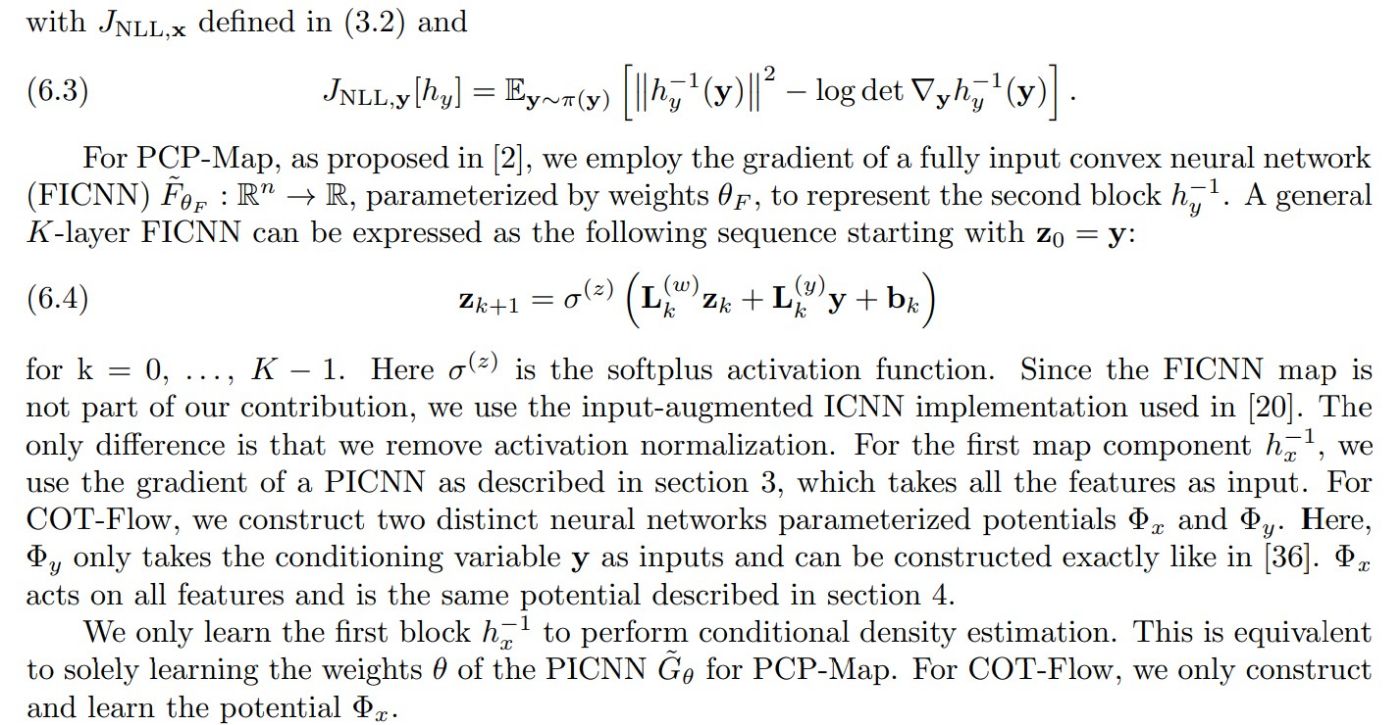
Since the reference density, ρz = N (0, In+m), is block-separable (i.e., it factorizes into the product ρz(zx, zy) = ρx(zx)ρy(zy)), we decouple the objective functional into the following two terms
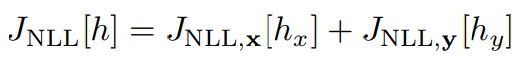
Table 3
Hyperparameter sample space for the UCI tabular datasets experiment.
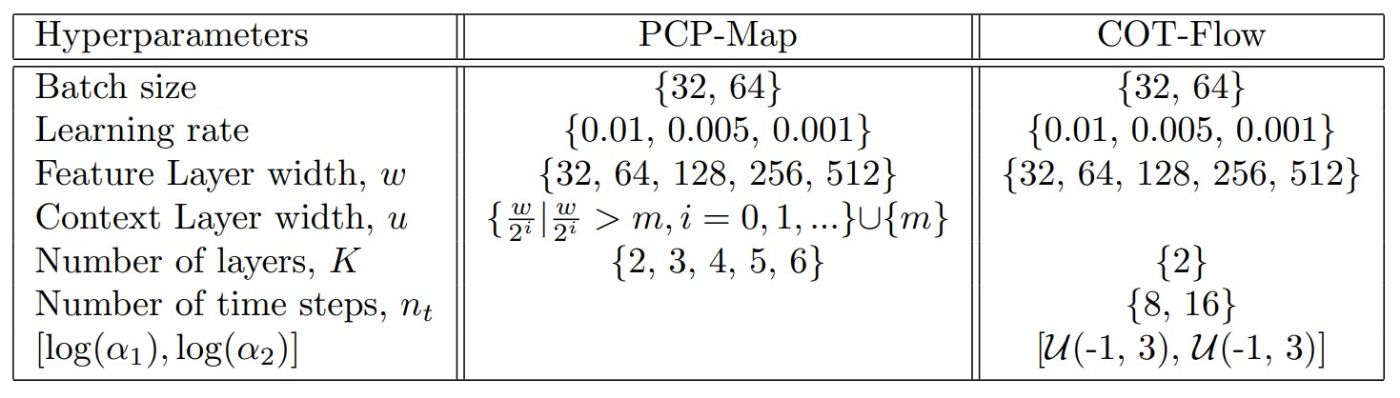


The hyperparameter sample space we use for this experiment is presented in Table 3. We select smaller batch sizes and large learning rates as this leads to fast convergence on these relatively simple problems. For each dataset, we select the ten best hyperparameter combinations based on a pilot run for full training. For PCP-Map’s pilot run, we performed 15 epochs for all three conditional datasets, three epochs for the Parkinson’s and the White Wine datasets, and four epochs for Red Wine using 100 randomly sampled combinations. For COT-Flow, we limit the pilot runs to only 50 sampled combinations due to a narrower sample space on model architecture. To assess the robustness of our approaches, we performed five full training runs with random initializations of network weights for each of the ten hyperparameter combinations for each dataset.
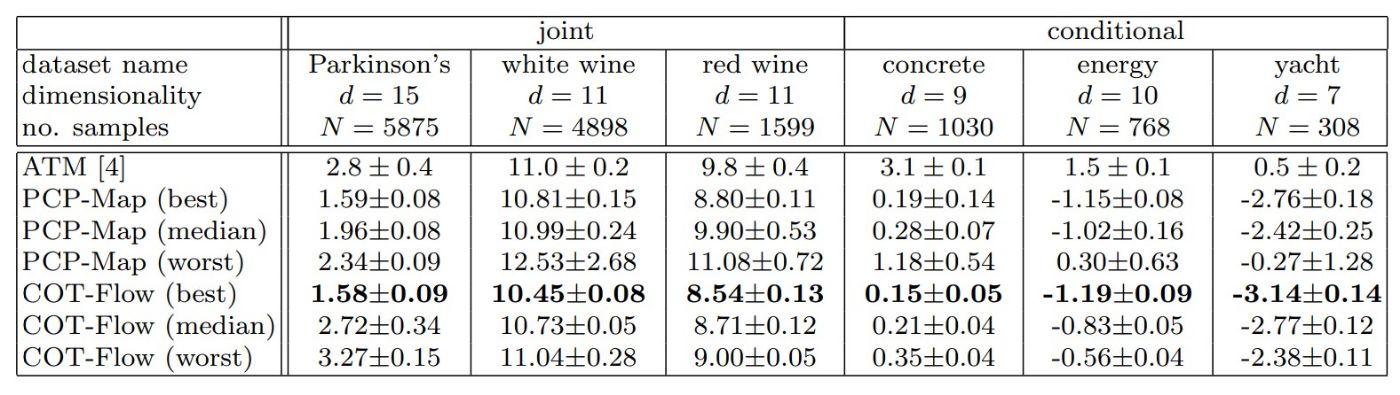
In Table 4, we report the best, median, and worst mean negative log-likelihood on the test data across the six datasets for the two proposed approaches and the best results for ATM. The table demonstrates that the best models outperform ATM for all datasets and that the median performance is typically superior. Overall, COT-Flow slightly outperforms PCP-Map in terms of loss values for the best models. The improvements are more pronounced for the conditional sampling tasks, where even the worst hyperparameters from PCP-Map and COT-Flow improve over ATM with a substantial margin.
Table 4
Mean negative log-likelihood comparisons between ATM, PCP-Map, and COT-Flow on test data. For our approaches, we report the best, median, and worst results over different hyperparameter combinations and five training runs. Lower is better, and we highlight the best results in bold.
6.2. Stochastic Lotka-Volterra.
We compare our approaches to an ABC approach based on Sequential Monte Carlo (SMC) for likelihood-free Bayesian inference using the stochastic LotkaVolterra (LV) model [55]. The LV model is a stochastic process whose dynamics describe the evolution of the populations S(t) = (S1(t), S2(t)) of two interacting species, e.g., predators and prey. These populations start from a fixed initial condition S(0) = (50, 100). The parameter x ∈ R 4 determines the rate of change of the populations over time, and the observation y ∈ R 9 contains summary statistics of the time series generated by the model. This results in an observation vector with nine entries: the mean, the log variance, the auto-correlation with lags 1 and 2, and the cross-correlation coefficient. The procedure for sampling a trajectory of the species populations is known as Gillespie’s algorithm.
Given a prior distribution for the parameter x, we aim to sample from the posterior distribution corresponding to an observation y∗. As in [37], we consider a log-uniform prior distribution for the parameters whose density (of each component) is given by π(log xi) = U(−5, 2). As a result of the stochasticity that enters non-linearly in the dynamics, the likelihood function is not available in closed form. Hence, this model is a popular benchmark for likelihood-free inference algorithms as they avoid evaluating π(y|x) [37].
We generate two training sets consisting of 50k and 500k samples from the joint distribution π(x, y) obtained using Gillespie’s algorithm. To account for the strict positivity of the parameter, which follows a log-uniform prior distribution, we perform a log transformation of the x samples. This ensures that the conditional distribution of interest has full support, which is needed to find a diffeomorphic map to a Gaussian. We split the log-transformed data into ten folds and use nine folds of the samples as training data and one fold as validation data. We normalize the training and validation sets using the training set’s empirical mean and standard deviation.
For the pilot run, we use the same sample space in Table 3 except expanding the batch size space to {64, 128, 256} for PCP-Map and {32, 64, 128, 256} for COT-Flow to account for the increase in sample size. We also fixed, for PCP-Map, w = u. During PCP-Map’s pilot run, we perform two training epochs with 100 hyperparameter combination samples using the 50k dataset. For COT Flow, we only perform one epoch of pilot training as it is empirically observed to be sufficient. We then use the best hyperparameter combinations to train our models on the 50k and 500k datasets to learn the posterior for the normalized parameter in the log-domain. After learning the maps, we used their inverses, the training data mean and standard deviation, and the log transformations to yield parameter samples in the original domain.
The SMC-ABC algorithm finds parameters that match the observations with respect to a selected distance function by gradually reducing a tolerance ϵ > 0, which leads to samples from the true posterior exactly as ϵ → 0. For our experiment, we allow ϵ to converge to 0.1 for the first true parameter and 0.15 for the second.
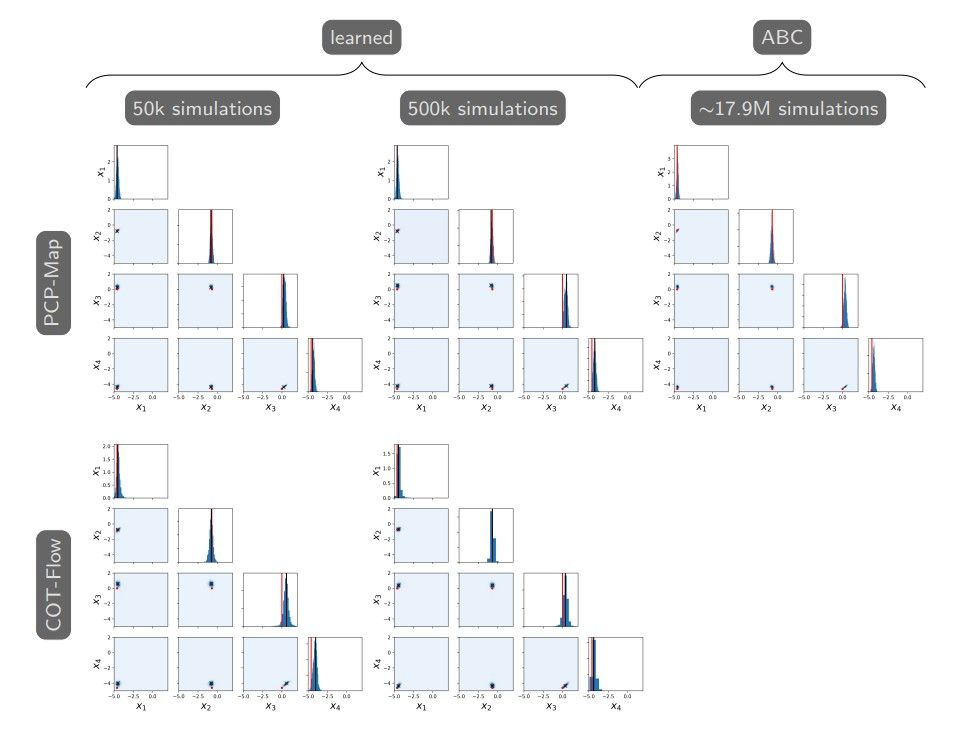
We evaluate our approaches for maximum-a-posteriori (MAP) estimation and posterior sampling. We consider a true parameter x∗ = (0.01, 0.5, 1, 0.01)⊤, which was chosen to give rise to oscillatory behavior in the population time series. Given one observation y∗ ∼ π(y|x ∗), we first identify the MAP point by maximizing the estimated log-likelihoods provided by our approaches. Then, we generate 2000 samples from the approximate posterior π(x|y∗) using our approaches. Figure 2 presents one and two-dimensional marginal histograms and scatter plots of the MAP point and samples, compared against 2000 samples generated by the SMC-ABC algorithm from [7]. Our
approaches yield samples tightly concentrated around the MAP points that are close to the true parameter x∗ .
Table 5
Sampling efficiency comparisons between PCP-Map and COT-Flow in terms of GPU time in seconds (s). We report the mean and standard deviation over five runs, respectively.
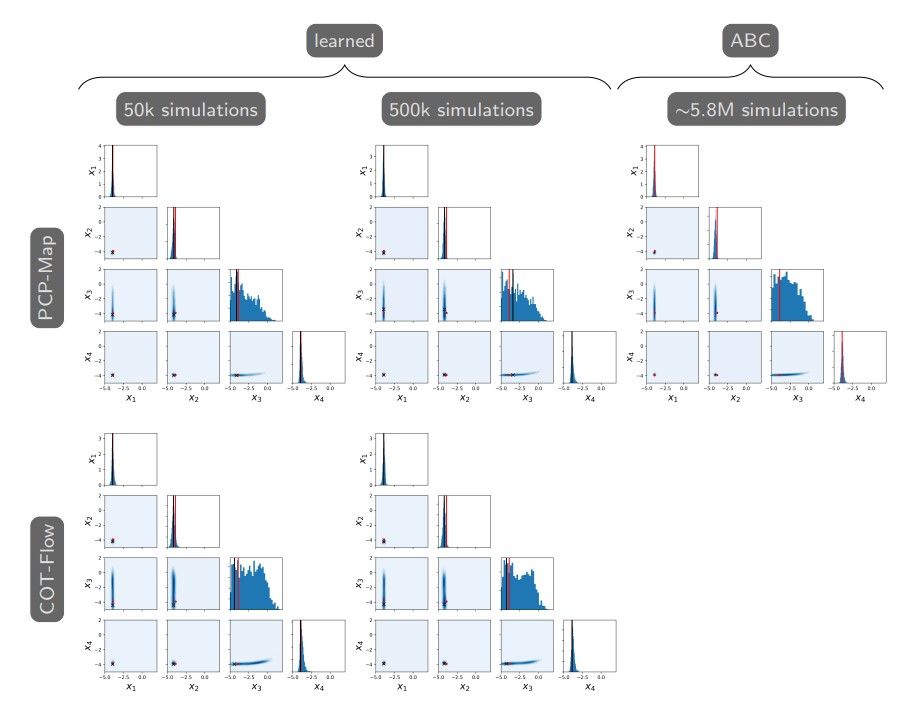
To provide more evidence that our learning approaches indeed solve the amortized problem, Figure 3 shows the MAP points and approximate posterior samples generated from a new random observation y∗ corresponding to the true parameter x∗ = (0.02, 0.02, 0.02, 0.02)⊤. We observe similar concentrations of the MAP point and posterior samples around the true parameter and similar correlations learned by the generative model and ABC, for example, between the third and other parameters.
Efficiency-wise, PCP-Map and COT-Flow yield similar approximations to ABC at a fraction of the computational cost of ABC. The latter requires approximately 5 or 18 million model simulations for each conditioning observation, while the learned approaches use the same 50 thousand simulations to amortize over the observation. These savings generally offset the hyperparameter search and training time for the proposed approaches, which is typically less than half an hour per full training run on a GPU. For some comparison, SMC-ABC took 15 days to reach ϵ = 0.1 for x∗ = (0.01, 0.5, 1, 0.01)⊤.
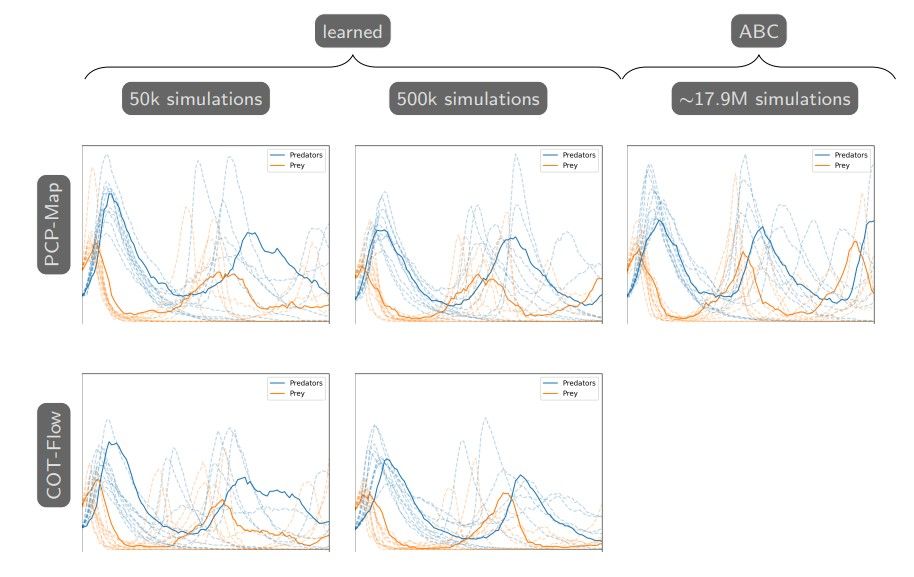
To further validate that we approximate the posterior distribution as well, Figure 4 compares the population time series generated from approximate posterior samples given by our approaches and SMC-ABC. This corresponds to comparing the posterior predictive distributions for the states S(t) given different posterior approximations for the parameters. While the simulations have inherent stochasticity, we observe that the generated posterior parameter samples from all approaches recover the expected oscillatory time series simulated from the true parameter x∗ , especially at earlier times.
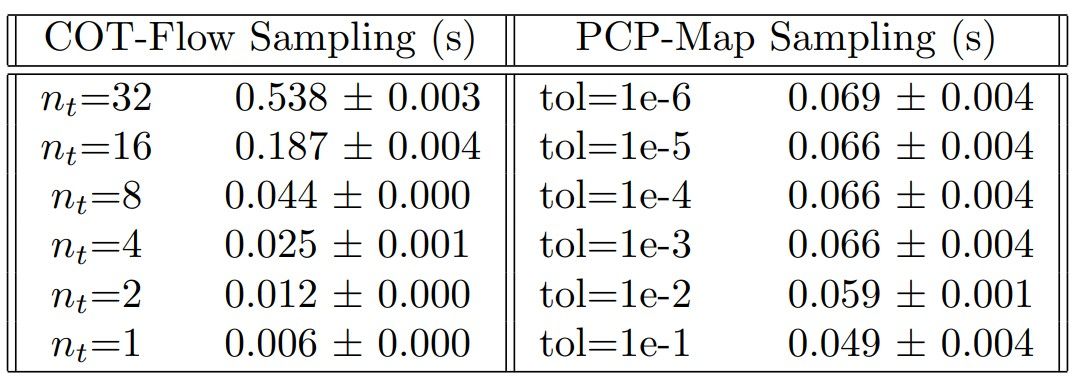
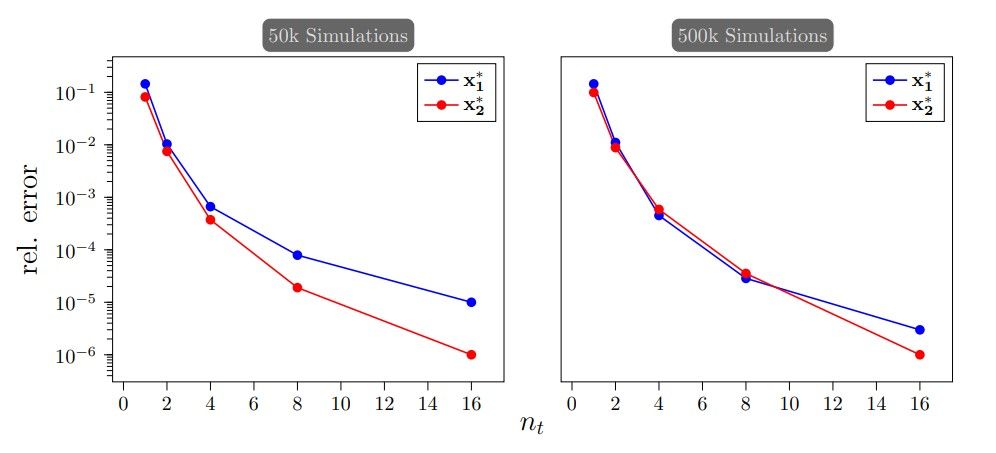
In the experiments above, we employed nt = 32 to generate posterior samples during testing for the COT-Flow. However, one can also decrease nt after training to generate samples faster without sacrificing much accuracy, as shown in Figure 5. Thereby, one can achieve faster sampling speed using COT-Flow than using PCP-Map as demonstrated in Table 5. To establish such a comparison, we increase the l-BFGS tolerance when sampling using PCP-Map, which produces a similar effect on sampling accuracy than decreasing the number of time steps for COT-Flow.
6.3. 1D Shallow Water Equations.
The shallow water equations model wave propagation through shallow basins described by the depth profiles parameter, x ∈ R 100, which is discretized at 100 equidistant points in space. After solving the equations over a time grid with 100 cells, the resulting wave amplitudes form the 10k dimensional raw observations. As in [40], we perform a 2D Fourier transform on the raw observations and concatenate the real and imaginary parts since the waves are periodic. We define this simulation-then-Fourier-transform process as our forward model and denote it as Ψ(x). Additive Gaussian noise is then introduced to the outputs of Ψ, which gives us the observations y = Ψ(x) + 0.25ϵ, where y ∈ R 200×100 and ϵi,j ∼ N (0, 1). We aim to use the proposed approaches to learn the posterior π(x|y).
We follow instructions from [40] to set up the experiment and obtain 100k samples from the
joint distribution π(x, y) as the training dataset using the provided scripts. We use the prior
distribution
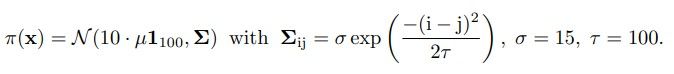
Using a principal component analysis (PCA), we analyze the intrinsic dimensions of x and y. To
ensure that the large additive noise 0.25ϵ does not affect our analysis, we first study another set of 100k noise-free prior predictives using the estimated covariance Cov(Y) ≈ 1 N−1Y⊤Y. Here, Y ∈ R 100000×20000 stores the samples row-wise in a matrix. The top 3500 modes explain around 96.5% of the variance. A similar analysis on the noise-present training dataset shows that the top 3500 modes, in this case, only explain around 75.6% of the variance due to the added noise. To address the rank deficiency, we construct a projection matrix Vproj using the top 3500 eigenvectors of Cov(Y) and obtain the projected observations yproj = V⊤ projy from the training datasets. We then perform a similar analysis for x and discovered that the top 14 modes explained around 99.9% of the variance. Hence, we obtain xproj ∈ R 14 as the projected parameters.
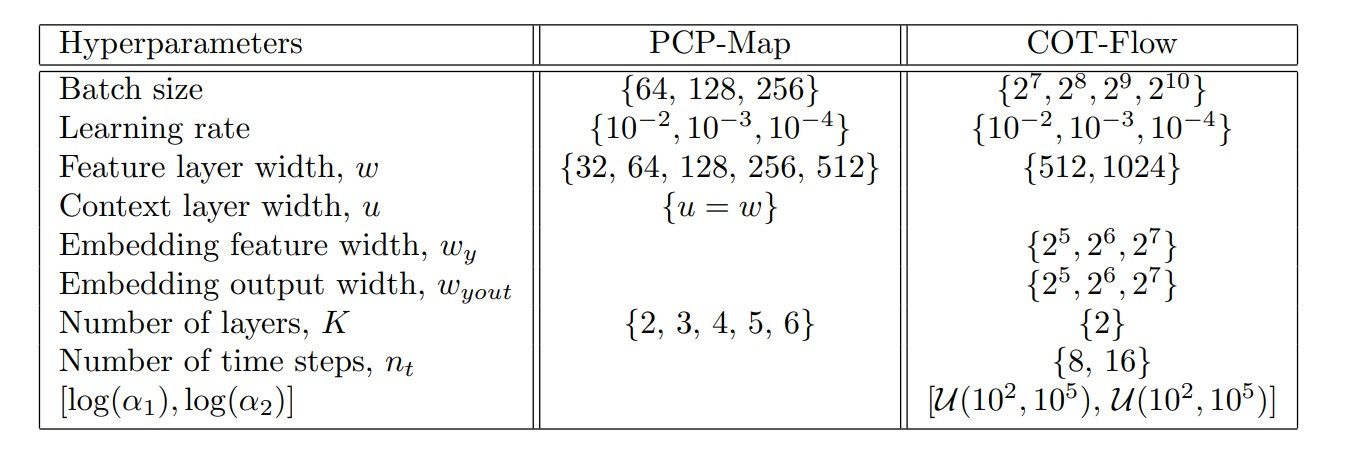
Table 6
Hyperparameter sample space for the 1D shallow water equations experiment.

We then trained our approaches to learn the reduced posterior, πproj(xproj|yproj). For comparisons, we trained the flow-based NPE approach to learn the same posterior exactly like did in [40]. For COT-Flow, we add a 3-layer fully connected neural network with tanh activation to embed yproj. To pre-process the training dataset, we randomly select 5% of the dataset as the validation set and use the rest as the training set. We project both x and y and then normalize them by subtracting the empirical mean and dividing by the empirical standard deviations of the training data.
We employ the sample space presented in Table 6 for the pilot runs. For COT-Flow, we select a w sample space with larger values for maximum expressiveness and allow multiple optimization steps over one batch, randomly selected from {8, 16}. We then use the best hyperparameter combination based on the validation loss for the full training. For NPE, we used the sbi package [50] and the scripts provided by [40].
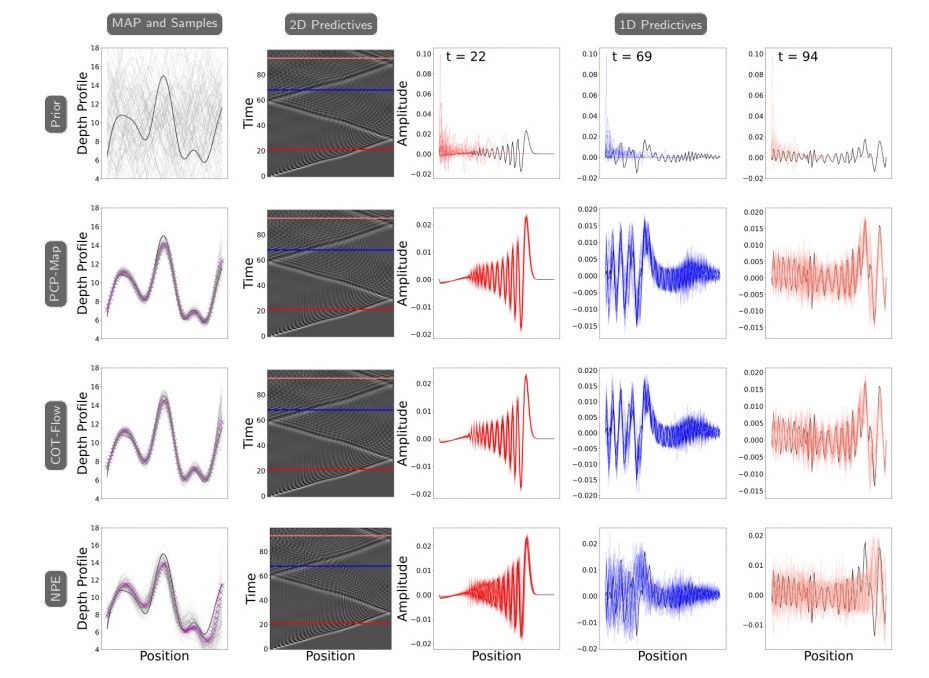
We first compare the accuracy of the MAP points, posterior samples, and posterior predictives across the three approaches. The MAP points are obtained using the same method as in subsection 6.2. For posterior sampling, we first sample a “ground truth” x∗ ∼ π(x) and obtain the associated ground truth reduced observation y∗ proj = V⊤ proj(Ψ(x∗) + 0.25ϵ). Then, we use the three approaches to sample from the posterior πproj(xproj|y ∗ proj). This allows us to obtain approximate samples x ∼ π(x|y∗). The posterior predictives are obtained by solving the forward model for the generated parameters. Through Figure 6, we observe that the MAP points, posterior samples, and predictives produced by PCP-Map and COT-Flow are more concentrated around the ground truth than those produced by NPE.
We perform the simulation-based calibration (SBC) analysis described in [40, App. D.2] to further assess the three approaches’ accuracy; see Figure 7. We can see that, while they are all well calibrated, the cumulative density functions of the rank statistics produced by PCP-Map align almost perfectly with the CDF of a uniform distribution besides a few outliers.
Finally, we analyze the three approaches’ efficiency in terms of the number of forward model evaluations. We train the models using the best hyperparameter combinations from the pilot run on two extra datasets with 50k and 20k samples. We compare the posterior samples’ mean and standard deviation against x∗ across three approaches trained on the 100k, 50k, and 20k sized datasets as presented in Figure 8. We see that PCP-Map and COT-Flow can generate posterior samples centered more closely around the ground truth than NPE using only 50k training samples, which translates to higher computational efficiency since fewer forward model evaluations are required.
6.4. Comparing PCP-Map to amortized CP Flow.
We conduct this comparative experiment using the shallow water equations problem for its high dimensionality. We include the more challenging task of learning π(x|yproj), obtained without projecting the parameter, to test the two approaches most effectively. We followed [20] and its associated GitHub repository as closely as possible to implement the amortized CP-Flow. To ensure as fair of a comparison as possible, we used the hyperparameter combination from the amortized CP-Flow pilot run for learning π(x|yproj) and the combination from the PCP-Map pilot run for learning πproj(xproj|yproj). Note that for learning π(x|yproj), we limited the learning rate to {0.0001} for which we observed reasonable convergence.
In the experiment, we observed that amortized CP-Flow’s Hessian vector product function gave NaNs consistently when computing the stochastic log-determinant estimation. Thus, we resorted to exact computation for the pilot and training runs. PCP-Map circumvents this as it uses exact log determinant computations. Each model was trained five times to capture possible variance.
For comparison, we use the exact mean negative log-likelihood as the metric for accuracy and
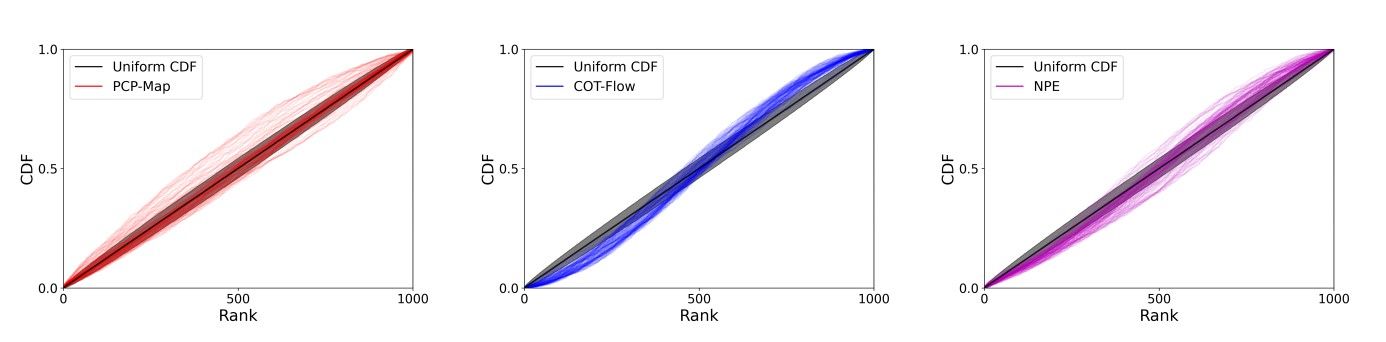
Table 7
Computational cost comparisons between amortized CP-Flow and PCP-Map in terms of GPU time in seconds (s). We report the mean and standard deviation of the training and validation mean NLL, the mean and standard deviation of the GPU times over five training runs, and the number of parameters in million(M), respectively.
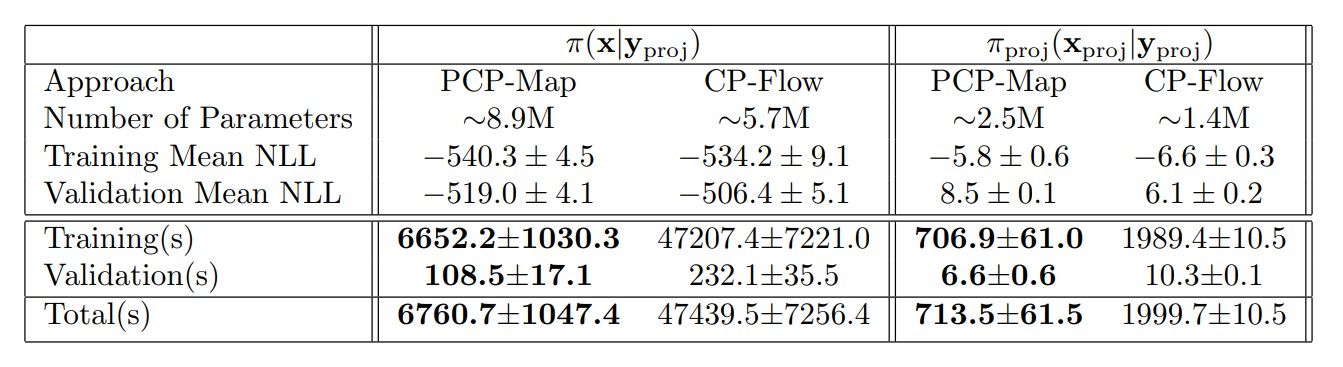
GPU time as the metric for computational cost. We only record the time for loss function evaluations and optimizer steps for both approaches. The comparative results are presented in Table 7. We see that PCP-Map and amortized CP-Flow reach similar training and validation accuracy. However, PCP-Map takes roughly 7 and 3 times less GPU time, respectively, on average, to achieve that accuracy than amortized CP-Flow. Possible reasons for the increased efficiency of PCP-Map’s are its use of ReLU non-negative projection, vectorized Hessian computation, removal of activation normalization, and gradient clipping.
